Count Objects in an Image with MXNet and Amazon SageMaker
Counting objects in images is one of the fundamental computer vision tasks that is easily handled by using Convolutional Neural Networks. In this tutorial, I am going to show how you can create a real-life application that accomplishes this task. This article is a hands-on end-to-end tutorial. We start from zero, and we will end up with a usable API. Here’s what we will be doing:
- Get acquainted with Amazon SageMaker.
- Generate training data.
- Build a CNN model.
- Train the model.
- Deploy the model.
- Test the model.
That is a lot to cover, so let us get started.
Amazon SageMaker
Amazon SageMaker is a service that provides the framework and the infrastructure to build, test and deploy machine learning enabled applications. I am going to assume you already have an AWS account and you are willing to spend a few dollars.
The typical workflow in Amazon SageMaker involves running a cheaper Jupyter Notebook instance for exploratory data analysis, data manipulation, and feature engineering. The next step is creating an Amazon SageMaker script. The main two tasks of this script are to train a model and to predict using the generated model.
Next comes the part where the actual training takes place using the Amazon SageMaker script. Training takes place on a more expensive machine.
After training completes successfully, the generated model is used along with the Amazon SageMaker script to enable a REST API endpoint that can be queried for predictions. A graphical overview of the process can be seen below, but it will become more evident as we advance.
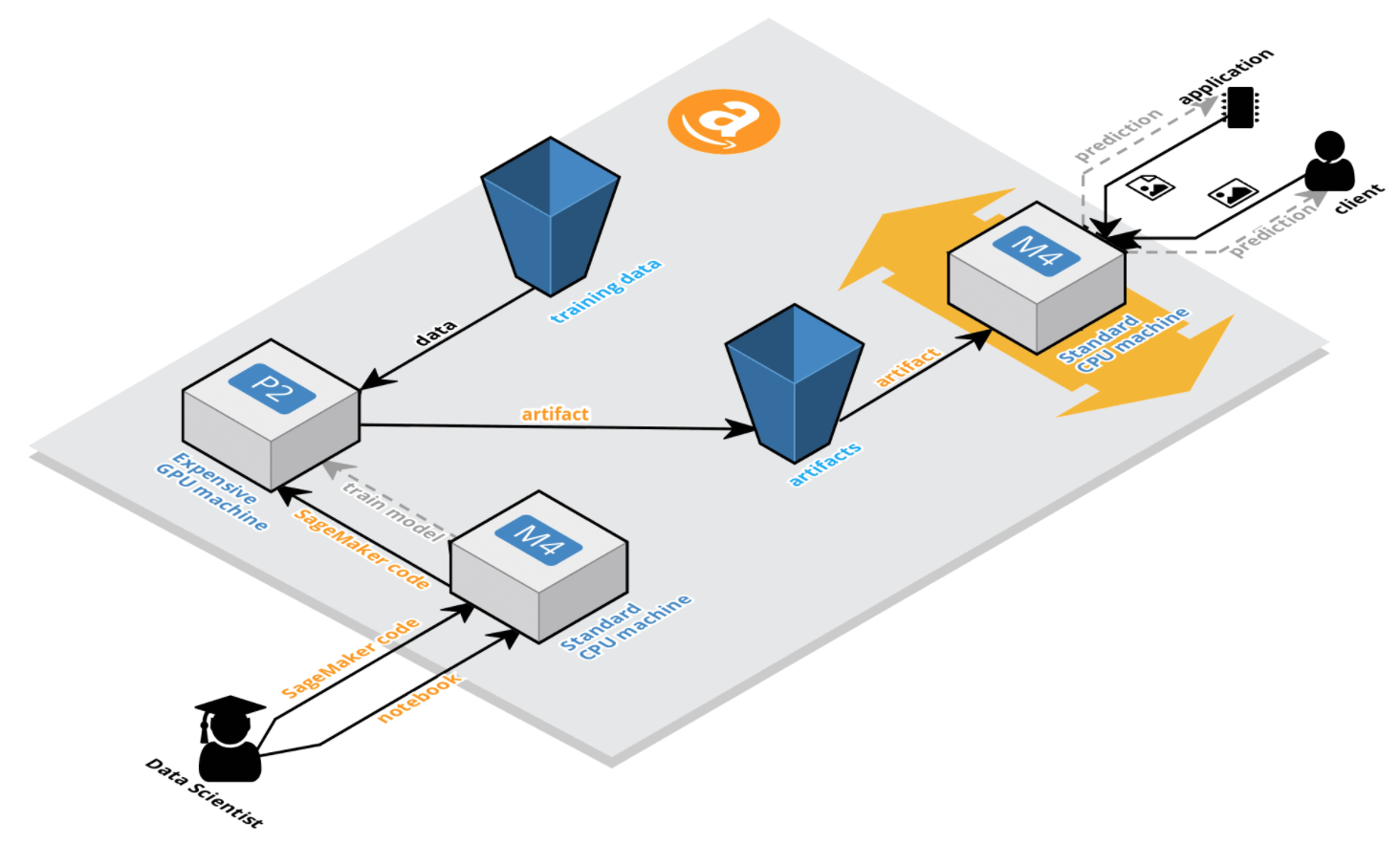
Starting a Notebook Instance
After logging in your AWS account, access the Amazon SageMaker console and create a Notebook Instance. For this tutorial, a cheaper ml.m4.xlarge would be appropriate.
Once the instance is operational, open Jupyter and create a new notebook based on the Conda MXNet Python 3 kernel.
Synthetic Data Generation
The data we are going to use in this tutorial will be generated from scratch, and it is going to be unique. No more wine tasting, MNIST, CIFAR or other overused datasets. It is probably good that different machine learning frameworks provide ready to use datasets, but at the same time, I feel that a lot of the many tutorials out there downplay the feature engineering part. In practice, data wrangling is necessary, takes a lot of time, and one cannot advance to the modeling part without having the data in the required format first. With that in mind, I am going to take you through the full process of data engineering all the way to the modeling part and beyond ![]() .
.
We will be generating synthetic data consisting of images that have various numbers of different shapes inside them. Here is an example of the type of images we are going to work with. These are typical RGB images stored as .png. There are three shapes: circle, square and triangle. We will try to count the number of circles, so that will be our target variable. Here are a few examples of the images we will be working with:

First, we define a set of parameters. You can change most of them freely.
# You can modify these ones
dataset_size = 12000
img_size = 128
shape_size = 8
max_rows = 14
max_cols = 14
target_shape = "circle"
# Do not modify these one
shapes = ["circle", "triangle", "square"]
img_width = img_height = img_size
The next part is about creating the source code files that are going to be fed in the Shaper compiler. As you can see from the code, the files have a random configuration. However, there is no guarantee that duplicates do not exist. For each generated file, we keep track of how many circles are in it, which is our target variable. The list of files, and the number of circles in them, are kept in a .csv file for easy handling via pandas.
import os
from random import randint, seed
# Make sure the required folder structure is in place
os.makedirs("data/raw")
# Make sure data generation is reproducible
seed(42)
with open("data/dataset.csv", "w+") as dataset:
# Write .csv header
dataset.write("img_path,target\n")
for i in range(1, dataset_size + 1):
# counter for the number of circles in the current image
cnt_target = 0
filename = str(i).zfill(6) + ".shape"
# Generate Shaper DSL source code. The first
# part is the same for all files
file_content = "img_dim:{},shp_dim:{}>>>" \
.format(img_size, shape_size)
# First we iterate on rows
for x in range(0, randint(1, max_rows)):
# Second we iterate on columns
for y in range(0, randint(1, max_cols)):
# Get the random shape
shape_idx = randint(0, len(shapes) - 1)
shape = shapes[shape_idx]
# If the shape is a circle update counter
if shape == target_shape:
cnt_target += 1
file_content = file_content + shape + ","
# Remove the last row separator
file_content = file_content[:-1]
# Add columns separator
file_content += "|"
# Remove the last columns separator
file_content = file_content[:-1]
# Add the termination sequence
file_content += "<<<"
with open("./data/raw/{}".format(filename), "w") as shape_file:
shape_file.write(file_content)
dataset.write("./raw/{}.png,{:d}\n".format(filename, cnt_target))
After running the above cell, you will end up with 12000 .shaper source code files. Also, the ./data/dateset.csv file contains the path of each of the images that will be generated in the next step along with their target variable (number of circles).
> head data/dataset.csv
img_path,target
./raw/000001.shape.png,30
./raw/000002.shape.png,23
./raw/000003.shape.png,24
./raw/000004.shape.png,1
./raw/000005.shape.png,16
./raw/000006.shape.png,33
./raw/000007.shape.png,8
./raw/000008.shape.png,38
./raw/000009.shape.png,22
The .shaper files contain source code that looks like this(except the newlines, the are no newlines in the .shaper files):
img_dim:128,
shp_dim:8
>>>
square,square|
square,square,circle|
triangle,triangle,triangle|
square,square,circle,square,triangle,circle,circle|
circle,square|
triangle,circle,square,triangle,circle,circle,square|
square,triangle,square,square,triangle,square,triangle,triangle,circle|
square,triangle,circle|
circle,circle,circle,square,triangle,triangle,square,triangle|
triangle,square,circle,square,square,circle,square,square,triangle|
square,triangle,circle,triangle,triangle,circle,circle
<<<
It is time to generate the images from the source code files. We will use a .jar binary for this purpose. In the Jupyter file browser, create a bin folder, and upload the .jar from here. For the security wary, the .jar can be generated by following the instructions in my Shaper DSL repository. Once the compiler is in its place, we can run the command to generate the .png files. It takes about a minute for 12000 files.
The reason for creating a DSL for generating images is that I did not want to depend on an existing dataset and also wanted to create training data at will, in the quantities that I would need. Another aspect is that while I could have just used JSON and a custom compiler, I wanted a language that had as few tokens as possible. My plan is to reverse engineer the images back to code, but this is something for another time ![]() .
.
import subprocess
subprocess.call("""
java -cp bin/shaper-all.jar \
com.cosminsanda.shaper.compiler.Shaper2Image \
--source-dir data/raw""", shell=True);
At this point, there will be 12000 .png images in the ./data/raw directory. These files represent our synthetic data.
Data Processing
We now have to split the data into training, validation, and test. The training dataset is the largest with 10000 samples while validation and test datasets have 1000 samples each. The split will be done at random. We will end up with three individual dataframes containing paths to images and their respective target variables.
from pandas import read_csv
import numpy as np
# Read the .cs
df = read_csv("data/dataset.csv")
train = df.sample(frac=.8333, random_state=42)
validation = df.loc[~df.index.isin(train.index), :] \
.sample(frac=.5, random_state=42)
# What this contraption does is that it selects all
# indexes that have not already been selected in the
# previous two dataframes
test = df.loc[np.logical_not(
np.logical_xor(
~df.index.isin(train.index),
~df.index.isin(validation.index))), :]
Next, I will define a function which takes an image and transforms it to an NDArray with which MXNet can work. The gist of it is that using OpenCV, we obtain the 128x128x3 numpy array representation of an RGB image. This array is then transformed to a 128x128x3 representation which is then normalized to 1. The target variable is also returned as part of the tuple.
import cv2
import mxnet as mx
def transform(row):
img = cv2.imread("./data/{}".format(row["img_path"]))
img = mx.nd.array(img)
img = img.astype(np.float32)
img = mx.nd.transpose(img, (2, 0, 1))
img = img / 255
label = np.float32(row["target"])
return img, label
Next, we are going to transform the training and validation datasets using the function above. We will end up with two lists of tuples. We are leaving the test dataset in place because we will use for testing in a real-life scenario, without much processing on the client side, so we want to keep the original images as they are.
train_nd = [transform(row) for _, row in train.iterrows()]
validation_nd = [transform(row) for _, row in validation.iterrows()]
To persist the training and validation datasets to disk, we are defining the save_to_disk function. We use Python’s pickle format because it is easy to save/load it.
import os
from pickle import dump
def save_to_disk(data, type):
os.makedirs("data/pickles/{}".format(type))
with open("data/pickles/{}/data.p".format(type), "wb") as out:
dump(data, out)
We are now going to use the function on the two datasets. After doing so, you will have two folders containing a file each, in the ./data/pickles/.
save_to_disk(train_nd, "train")
save_to_disk(validation_nd, "validation")
Here are the files:
> tree -h data/pickles
data/pickles
├── [ 96] train
│ └── [1.8G] data.p
└── [ 96] validation
└── [188M] data.p
Next, we need to initialize an Amazon SageMaker session. The client will make it easy for us to upload data to Amazon S3 and also return arguments that will be required in training further on.
import sagemaker
sagemaker_session = sagemaker.Session()
Using the SageMaker client, we will upload the two pickles to S3 in a location of your choice. I have redacted the details of my account, so feel free to modify as needed so that it matches your setup.
inputs = sagemaker_session.upload_data(path="data/pickles",
bucket="redacted",
key_prefix="sagemaker/demo")
As for the test dataset, we will copy the images to a ./test folder which is partitioned based on the number of circles in images.
from shutil import rmtree, copy2
# Cleanup first
rmtree("./test", True)
os.makedirs("./test")
for _, row in test.iterrows():
os.makedirs("test/{}".format(row["target"]), exist_ok=True)
copy2("./data/{}".format(row["img_path"]),
"./test/{}".format(row["target"]))
The file structure now looks like this:
> ls test
0 1 10 11 12 13 14 15 16 17 18 19 2 20
21 22 23 24 25 26 27 28 29 3 30 31 32 33
34 35 36 37 38 39 4 40 41 42 43 44 45 46
47 49 5 50 6 7 8 9
The last step in the data processing part is just to remove the ./data directory. We no longer need it.
rmtree("data", True)
Model Training
So this is where it gets interesting. Training on Amazon SageMaker means that a new powerful (and potentially expensive) machine is launched. The machine copies the needed data (training and validation) from S3, and then uses a script which conforms to an interface to execute training and saving the outputted model. For the case of MXNet, the script should conform to the documentation for the MXNet Estimator.
MXNet and Gluon
MXNet is a deep learning framework developed by the same folks behind the extraordinary XGBoost. It is now the defacto deep learning framework at Amazon and is part of the Apache incubator. DMLC, the group from which it originates, focuses on performance and distributed learning. MXNet has a higher level API, Gluon, which is actually what we will be using.
Amazon SageMaker Script - Part I - Training
At this point, you need to go back to your Jupyter file browser (do not close the opened notebook, you will need it later). Create a new Python file and name it object-counting-sagemaker-script.py. While adhering to the guidelines in the documentation, we will create the following functions:
def train(hyperparameters, channel_input_dirs, num_gpus):
pass
def save(net, model_dir):
pass
train function
Hyperparameters are going to be sent from the Jupyter notebook and allow us to configure arbitrary arguments that can be used by the training function. For our case, we will configure the batch_size and the number of epochs. Defaults for the two settings are also provided.
batch_size = hyperparameters.get("batch_size", 64)
epochs = hyperparameters.get("epochs", 3)
The channel_input_dirs is used by Amazon SageMaker framework to provide the location on disk where the data from S3 is copied to (remember what we uploaded to S3 earlier?). We load the pickles, and so we have the same variables in place as in the Jupyter notebook.
mx.random.seed(42)
training_dir = channel_input_dirs['training']
logging.info("Loading data from {}".format(training_dir))
with open("{}/train/data.p".format(training_dir), "rb") as pickle:
train_nd = load(pickle)
with open("{}/validation/data.p".format(training_dir), "rb") as pickle:
validation_nd = load(pickle)
Data is provided during training using Gluon’s DataLoader which is a type of iterator for batches.
train_data = DataLoader(train_nd, batch_size, shuffle=True)
validation_data = DataLoader(validation_nd, batch_size, shuffle=True)
The model
The architecture of the neural network follows., and as you can see, it is a straightforward setup around two convolutional network layers. Nothing fancy, but as you will see, it delivers excellent results in just a few epochs.
The with net.name_scope() is explained here.
Notice how we do not need to specify the input shape of the data. This is a very handy feature. The last layer is activated linearly which fits our purpose. Our problem is sort of a regression with discrete numbers rather than continuous values. This is similar to the problems that can be solved by a ordered logistic regression algorithm.
net = Sequential()
with net.name_scope():
net.add(Conv2D(channels=32, kernel_size=(3, 3),
padding=0, activation="relu"))
net.add(Conv2D(channels=32, kernel_size=(3, 3),
padding=0, activation="relu"))
net.add(MaxPool2D(pool_size=(2, 2)))
net.add(Dropout(.25))
net.add(Flatten())
net.add(Dense(1))
MXNet works either in the context of CPU or GPU, based on the number of GPUs that are available (the num_gpus argument to the function) we use either. The parameters of the network are initialized using the Xavier initializer (a.k.a Glorot). This gives noticeable better performance faster. The loss will be the mean squared error. For optimizer, we use the popular adam, although I hope we can soon use the new optimizers from MXNet version 1.1.
ctx = mx.gpu() if num_gpus > 0 else mx.cpu()
net.collect_params().initialize(Xavier(magnitude=2.24), ctx=ctx)
loss = L2Loss()
trainer = Trainer(net.collect_params(), optimizer="adam")
Now, the actual training is not very convenient. Unfortunately, you have to call forward propagation and back propagation manually. The data used in training is made available on the GPU, that is the reason for the as_in_context. A custom function for measuring the performance in counting the number of shapes is also referenced, calc_perf (the implementation is in the next code block). In the end, the function returns the Gluon model object, net.
smoothing_constant = .01
for e in range(epochs):
moving_loss = 0
for i, (data, label) in enumerate(train_data):
data = data.as_in_context(ctx)
label = label.as_in_context(ctx)
with autograd.record():
output = net(data)
loss_result = loss(output, label)
loss_result.backward()
trainer.step(batch_size)
curr_loss = nd.mean(loss_result).asscalar()
if (i == 0) and (e == 0):
moving_loss = curr_loss
else:
moving_loss = (1 - smoothing_constant) * moving_loss + \
smoothing_constant * curr_loss
trn_total, trn_detected = calc_perf(net, ctx, train_data)
val_total, val_detected = calc_perf(net, ctx, validation_data)
log = "Epoch {} loss: {:0.4f} perf_test: {:0.2f} perf_val: {:0.2f}" \
.format(e, moving_loss,
trn_detected / trn_total,
val_detected / val_total)
logging.info(log)
return net
The performance function just uses the current epoch’s model to make predictions on the validation set. It then uses the rounded results to see how many images have had their target value predicted correctly. It returns a tuple with the total number of images analyzed and the total number of images for which we had a correct prediction.
def calc_perf(model, ctx, data_iter):
raw_predictions = np.array([])
rounded_predictions = np.array([])
actual_labels = np.array([])
for i, (data, label) in enumerate(data_iter):
data = data.as_in_context(ctx)
label = label.as_in_context(ctx)
output = model(data)
predictions = nd.round(output)
raw_predictions = np.append(raw_predictions,
output.asnumpy().squeeze())
rounded_predictions = np.append(rounded_predictions,
predictions.asnumpy().squeeze())
actual_labels = np.append(actual_labels,
label.asnumpy().squeeze())
results = np.concatenate((raw_predictions.reshape((-1, 1)),
rounded_predictions.reshape((-1, 1)),
actual_labels.reshape((-1, 1))), axis=1)
detected = 0
i = -1
for i in range(int(results.size / 3)):
if results[i][1] == results[i][2]:
detected += 1
return i + 1, detected
save function
The save function is used for serializing a model. The function receives as argument the output of the train function, that is the model object. The model object is then serialized to two files:
- A
.jsonfile which stores the architecture of the neural network. - A
.paramsfile which stores the parameters of the neural network.
def save(net, model_dir):
y = net(mx.sym.var("data"))
y.save("{}/model.json".format(model_dir))
net.collect_params().save("{}/model.params".format(model_dir))
After this function gets executed, the serialized model gets packed into an artifact and then uploaded to Amazon S3.
The Amazon SageMaker script file should now look similar to this gist (opens in a new tab).
Initialising Training
Back to the Jupyter notebook, we will start training. We create an Amazon SageMaker estimator, and we configure some parameters. First, we provide the filename of the training script. Next, we need the role for execution, which you should be able to get automatically if running on the notebook instance. We will use a single GPU (more expensive) machine to do the training. We will train for 5 epochs and use batches of 64 samples. Lastly, training will execute in a Python 3 environment.
estimator = sagemaker.mxnet.MXNet("object-counting-sagemaker-script.py",
role=sagemaker.get_execution_role(),
train_instance_count=1,
train_instance_type="ml.p2.xlarge",
hyperparameters={"epochs": 5},
py_version="py3")
We now trigger the training execution and wait for about 10 minutes. The input to the fit function is the S3 path where the train and validation data was uploaded. This information is used automatically by the framework for making the data available on the training machine.
estimator.fit(inputs)
The notebook outputs some information, but it gets interesting towards the last part. Here is a sample:
Epoch :0 loss: 56.1768 perf_test: 0.20 perf_val: 0.21
Epoch :1 loss: 0.7820 perf_test: 0.61 perf_val: 0.59
Epoch :2 loss: 0.1410 perf_test: 0.90 perf_val: 0.87
Epoch :3 loss: 0.1065 perf_test: 0.89 perf_val: 0.90
Epoch :4 loss: 0.0920 perf_test: 0.88 perf_val: 0.86
===== Job Complete =====
Billable seconds: 324
This shows us how training performs. Also, you can see how I was billed for 324 seconds, which is about 5 minutes, so approximately 10 cents. The cool thing is that I paid for using GPU power in an optimized fashion. I have not used the GPU for data processing which I already did, and it also has not idled.
At this point, there should be an artifact on S3 containing the serialized model.
Amazon SageMaker Script - Part II - Deployment
It is now time to put our model to good use by creating a real-life application endpoint that can be queried online. The way deployment works is that a new machine is launched which acts as a REST endpoint. The API it exposes can then be used to make inferences. The endpoint machine has a copy of the artifact generated when training. It uses the Amazon SageMaker script to load the embedded model.
Since we are using a Gluon model we need to specify in the Amazon SageMaker script a few things:
- How the model is loaded.
- What are the transformations applied to the incoming data.
- How is prediction executed.
- How is the data returned to the client.
To do all of this, we go back to the object-counting-sagemaker-script.py file. We need to create the following functions:
def model_fn(model_dir):
pass
def transform_fn(model, input_data, content_type, accept):
pass
model_fn function
The model_fn function is used by the Amazon SageMaker framework to load a model from an artifact. The model_dir represents where on the local disk the artifact has been extracted. In our case, the Gluon model can be loaded like this:
def model_fn(model_dir):
with open("{}/model.json".format(model_dir), "r") as model_file:
model_json = model_file.read()
outputs = mx.sym.load_json(model_json)
inputs = mx.sym.var("data")
param_dict = gluon.ParameterDict("model_")
net = gluon.SymbolBlock(outputs, inputs, param_dict)
# We will serve the model on CPU
net.load_params("{}/model.params".format(model_dir), ctx=mx.cpu())
return net
Recreating the model object from its serialized form is pretty standard. I think the only aspect that needs explaining is the ctx argument in the load_params function. I have explicitly set the context to match the machine I will be using for serving the model. That is, I will be using a CPU based machine for hosting the model. Have I used a GPU machine, I would have initiated the context to mx.gpu().
The function returns the Gluon model object.
transform_fn function
This is a do it all kind of a function, where we receive data, we engineer it, use it for prediction and then return the results. The Amazon SageMaker platform supposedly supports these functionalities as separate individual functions, but I could not make them work like that. On the same note, I find debugging in Amazon SageMaker somewhat tricky. I am pretty sure this will change in the future for the better, but for now, take it as a warning.
One of the arguments to the transform_fn function, is the content_type of the incoming data from the client. Our transform function will be nice enough to handle both batch and individual predictions. To that extent, when we receive content as application/png we will consider it to be a single instance. Similarly, when we receive content as application/json we will consider it batch.
Before I illustrate the transform_fn function, I have a helper function img2arr that is being used to transform incoming data to the format that MXNet can easily work with. Let me break it down for you:
- The PNG files that are going to be the input to the service need to be sent base64 encoded, so the first step in the helper function is to decode them.
base64is nice common denominator for sending binary data over the wire to REST endpoints. - The binary image then undergoes a transformation that finally results in the
numpyrepresentation of the image. - The
numpyarray is then transformed to anNDArraywhich gives us thetransposefunction. - The shape of inputs with which the convolutional networks in the model work is slightly different from the way the image is represented by default, so we need to reshape the data accordingly so that we end up with a 3x128x128 as opposed to 128x128x3. The axis are switched.
- The values in the array are then normalized to 1.
- The model is supposed to work with batches, so next we reshape the data as if it is a batch of just one element, so now we have 1x3x128x128.
- Lastly, we convert the
NDArrayback to anumpyarray. This will allow for further transformation when working in batch mode.
def img2arr(base64img):
img = base64.b64decode(base64img)
img = np.asarray(bytearray(img), dtype=np.uint8)
img = cv2.imdecode(img, cv2.IMREAD_COLOR)
img = img.astype(np.float32)
img = mx.nd.array(img)
img = mx.nd.transpose(img, (2, 0, 1))
img = img / 255
img = img.reshape((1, 3, 128, 128))
img = img.asnumpy()
return img
On to the transform_fn function, here is the implementation for the case when we just want one prediction. The image representation from the img2arr function is used directly in the model. The response is transformed from an NDArray to numpy and then flattered with ravel. Lastly, it is cast to a list so that json.dumps can work easily with it. The function returns a tuple, where the second part is the accept content type.
def transform_fn(model, input_data, content_type, accept):
if content_type == "application/png":
img = nd.array(img2arr(input_data))
response = model(img).asnumpy().ravel().tolist()
return json.dumps(response), accept
else:
raise ValueError("Cannot decode input to the prediction.")
When working with batch, there is slightly more to do. First, the input is considered to be an array of base64 encoded images. All of the images are then transformed with the img2arr function and collected as a list. The list is then used by the concatenate function to create a large batch. That means that if we have received 42 images from the client, we will end up with a matrix with the following dimensions: 42x3x128x128. The array is then transformed to an NDArray, and finally, it is used directly by the model to make predictions. The predictions then go through the same transformations as for the single image prediction.
def transform_fn(model, input_data, content_type, accept):
if content_type == "application/png":
img = nd.array(img2arr(input_data))
response = model(img).asnumpy().ravel().tolist()
return json.dumps(response), accept
elif content_type == "application/json":
json_array = json.loads(input_data, encoding="utf-8")
imgs = [img2arr(base64img) for base64img in json_array]
imgs = np.concatenate(imgs)
imgs = nd.array(imgs)
response = model(imgs)
response = nd.round(response)
response = response.asnumpy()
response = response.ravel()
response = response.tolist()
return json.dumps(response), accept
else:
raise ValueError("Cannot decode input to the prediction.")
The complete listing of the object-counting-sagemaker-script.py file can be found in this gist.
Now, ideally, both the modeling and the deploying functions would have been written at once. I say, ideally, because the model artifact resulted from training is then used for deployment as well, you cannot edit it straightforwardly. The problem is that the hosting part of the script (transform_fn and model_fn) need to be there already. I mean, you could go in S3 and replace the artifact with another one, but that would be inconvenient. A much better alternative is to just update the object-counting-sagemaker-script.py with the complete listing and execute training again. So back in Jupyter you would need to execute the estimator.fit(inputs) cell again. The model will then go through training again, and a new artifact will be created. The new artifact contains the hosting part so let’s go ahead and deploy it.
predictor = estimator.deploy(1, "ml.m4.xlarge")
Like I have mentioned before, we deploy to a CPU instance. That is it, when deployment finishes (takes a few minutes) we will have an endpoint which can be used for making predictions.
Testing the Endpoint
We have worked a lot to get to this point, let us see if the application we have built works. For this purpose, we will use the test dataset of images we have set aside.
First, create an Amazon SageMaker runtime client. This is used to call the endpoint easily.
import boto3
sagemaker_runtime_client = boto3.client("sagemaker-runtime")
Next, we will loop through each of the folders in the ./test folder. Remember, each folder holds images that contain the number of circles corresponding to the name of the said folder. We create batches and send them to the endpoint for inference. One thing to note is that you cannot have a payload heavier than 5MB. Each image file is base64 encoded, just like I have mentioned earlier. Then the batch is an array of batch64 encoded images. The batch is then binary encoded, and the request is made to the endpoint. We get back the inferences (rounded to the nearest integer). With these results, we can assess how good is our model at counting the circles in the images.
for dir in os.listdir("./test"):
batch = []
for file in os.listdir("./test/{}".format(dir)):
with open("./test/{}/{}".format(dir, file), "rb") as image_file:
# Decode the bas64 encoded image from its binary form
encoded_img = base64.b64encode(image_file.read()) \
.decode("utf-8")
# Add the image to the batch
batch.append(encoded_img)
# Create the binary payload
binary_json = json.dumps(batch).encode("utf-8")
response = sagemaker_runtime_client.invoke_endpoint(
EndpointName=predictor.endpoint,
Body=binary_json,
ContentType="application/json",
Accept="application/json"
)["Body"].read()
# The response is a JSON array
individual_predictions = json.loads(response, encoding="utf-8")
# Check if the prediction matches the directory number
# from which the batch was created
total = 0
detected = 0
for prediction in individual_predictions:
total += 1
if int(prediction) == int(directory):
detected += 1
# Print results for each directory
print("""Images with {} circles:
Total: {}
Detected: {}
Accuracy: {:0.2f}
""".format(directory, str(total), str(detected), detected/total))
The output will look something like this:
Images with 50 circles:
Total: 2
Detected: 0
Accuracy: 0.00
Images with 32 circles:
Total: 23
Detected: 13
Accuracy: 0.57
...
That is it. Finally, we are at the end, but now we have a real-life application that can count circles in an image. The full listing for the notebook in Amazon SageMaker can be found in this gist.
A sample GitHub respository is also available.
End Notes
About the Application
While the application we have just built is as real as anything else, it should be noticed that I have not tested it with images other than the ones I generate myself. This is something that can obviously be done, say try to use it against hand-drawn images.
There are few to no safeguards in place when using the application. For example, you need to use it with images that are 128x128x3. This can easily be parameterized, and the fact that the Gluon model does not require the input shape can be of great help.
About Amazon SageMaker
I think Amazon SageMaker is a great tool that formalizes Machine Learning and encourages best practices. There are still some rough edges, but on the other hand, it is a new service after all. I think some focus should be put on faster iterations and perhaps easier debugging when building the models and deploying the models.
This article presents just one way of using Amazon SageMaker. Another alternative is to just run the notebook on your machine.

Comments